Wrangling proteins gone wild
proteins sometimes run amuck. All the good stuff (the useful genetic and biological material) they contain can get distorted. Mutations in specific amino acids can cause long strands of proteins to curl in on themselves (like a ball of wool a cat has played with) and refuse to break apart. These strands, known as amyloid fibrils, can be extremely toxic and are usually harmful. They attach to organs like the brain and pancreas, preventing them from functioning as they should. They are responsible for diseases as seemingly different as diabetes and Alzheimer's, to name just a couple. Developing effective medications to treat these diseases, and cause the fibrils to dissolve typically involves biochemists in a lengthy and expensive process of trial and error.
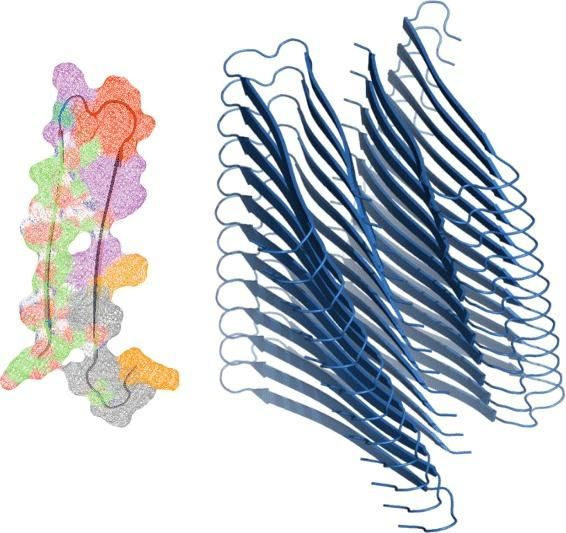
This is an amyloid fibril made up of 25 mis-folded amyloid proteins. (Right) A single color-coded amyloid protein. The different colors represent key stability regions explored by the algorithm developed at McGill. (Left)
Jerome Waldispuhl + Mohamed Smaoui
Billions of choices
But now McGill researchers, led by Prof. Jérôme Waldispühl of the School of Computer Science, have created a suite of computer programs that should speed up the process of drug discovery for diseases of this kind. The programs are designed to scan the fibrils (or misfolded proteins) looking for weak spots. The idea is to then design helpful genetic mutations to dissolve the bonds that hold the fibrils together - a bit like finding the right strand of wool to tug on to unravel a whole knotted ball. It's potentially a gargantuan task, because looking for the mutations that will prove useful in drug development involves exploring millions of possible structural combinations of genetic material.
But for the Fibrilizer, as McGill has dubbed its suite of computer tools, a name that hints at the super heroic nature of the programs they have developed, the task is of a very different order. "Within the space of a week, by using our programs and a supercomputer, we were able to look at billions of possible ways to weaken the bonds within these toxic protein strands. We narrowed it down to just 30 - 50 possibilities that can now be explored further," says Mohamed Smaoui, a McGill postdoctoral fellow and the first author on three recent papers on the research. "Typically biochemists can spend months or years in the lab trying to pinpoint these promising mutations."
Supercomputing to the rescue
The researchers tested their program on a medical compound that scientists have been trying to improve for the last couple decades. The compound is administered as part of a drug that is used by diabetes patients to boost the performance of insulin and is sold under the name Symlin. The synthetic compound is based on a version of the protein amylin, yet is known to be toxic to the pancreas over the long-term, creating amyloid fibrils. The McGill team were able to use Fibrilizer to pinpoint a limited number of possible genetic modifications to the compound that would act to reduce its toxicity.
Jérôme Waldispühl, the lead researcher on the papers, believes that computational research of this kind will play an increasingly important role in drug discovery in the future. "Computers are transforming the way that drugs are being developed," says Waldispühl. "Amyloid research has accelerated in the last 10 years. But computers may prove to be the key to finding better medications for a whole range of systemic and neurodegenerative diseases, from arthritis to Parkinson's. Without supercomputers and programs of this kind, it would take much longer and be much more expensive to do this kind of research and come up with these possible solutions to the problem."