Microchip allows optical observation of single molecules
Cornell scientists create microchip with light-impeding holes for detailed, optical observation of single molecules in their natural state
ITHACA, N.Y. -- Using a novel technique, supported largely by off-the-shelf instruments, scientists at Cornell University have for the first time optically isolated individual biological molecules in naturally occurring molecular concentrations and watched their complex behavior as they interact with a protein.
The technique, made possible by the ability of nanofabrication to produce a microchip with light-impeding holes with a diameter one-tenth of the wavelength of light, could promise a new method of DNA sequencing by which the genetic code can be "read" from a single DNA molecule.
It also promises to aid in future drug discovery because "it provides a very powerful way of looking at fluctuations and variability in behavior of individual enzyme molecules. We are seeing those variations, and they are huge," says Watt Webb, professor of applied and engineering physics at Cornell. "Observing them with such detail was hardly accessible until this experiment."
His report on watching individual molecules at work, "Zero-Mode Waveguides for Single-Molecule Analysis at High Concentrations," appears in the latest issue of the journal Science (Jan. 31, 2003). The article, which is illustrated on the cover of Science , also is authored by a multidisciplinary group of Cornell researchers: Michael Levene, an optics specialist and postdoctoral associate in applied and engineering physics; Jonas Korlach, a biologist who is a graduate student in biochemistry, molecular and cell biology; former postdoctoral associate Stephen Turner, now president and chief scientific officer of Nanofluidics, a Cornell spinoff; applied physics graduate student Mathieu Foquet: and applied and engineering physics professor Harold Craighead.
"This is an example of the possibilities provided by integrating nanostructures with biomolecules," says Craighead, who also is co-director of the National Science Foundation (NSF)-funded Nanobiotechnology Center at Cornell. "It represents a major step in the ability to isolate a single active biomolecule for study. This can be extended to other biological systems."
Until now, researchers were constrained from seeing individual molecules of an enzyme (a complex protein) interacting with other molecules under a microscope at relatively high physiological concentrations -- their natural environment -- by the wavelength of light, which limits the smallest volume of a sample that can be observed. This, in turn, limits the lowest number of molecules that can be observed in the microscope's focal spot to more than 1,000. Internal reflection microscopes have managed to reduce the number of molecules to about 100. But because this number is still far too high to detect individual molecules, significant dilution of samples is required.
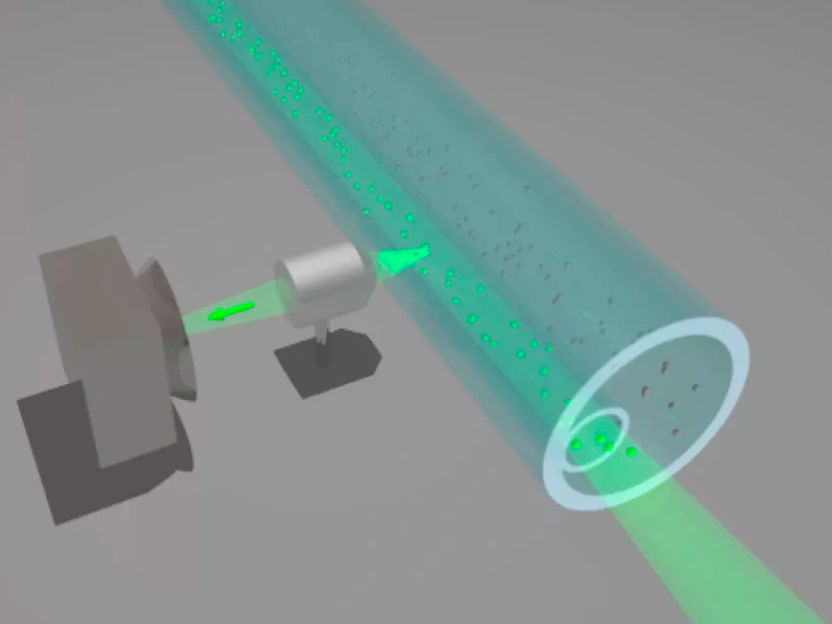
The researchers have discovered a way around these limitations, and in the process reduced the sample being observed 10,000-fold to just 2,500 cubic nanometers (1 nanometer is the width of 10 hydrogen atoms, or 1 billionth of a meter), by creating a microchip that actually prevents light from passing through and illuminating the bulk of the sample. The microchip, engineered from aluminum and glass in the Cornell Nanoscale Science and Technology Facility, a NSF-funded national center, contains 2 million holes (each called a waveguide), some as tiny as 40 nanometers in diameter, or one-tenth of the wavelength of light.
Small droplets of a mixture containing enzymes and specially prepared molecules were pipetted into wells on the microchip and placed the chip in an optical microscope. Each of the chip's holes is so tiny that light from a laser beam is unable to pass through and instead is reflected by the microchip's aluminum surface, with some photons "leaking" a short distance into the hole, on the bottom of which an enzyme molecule is located.
These few leaking photons are enough to illuminate fluorescent molecules, called fluorophores, attached as "tags" to nucleotides (molecules that make up the long chains of DNA) in the sample. In this way, the researchers were able to observe, for the first time, the interaction between the ligand (the tagged nucleotide) and the enzyme in the observation volume (the region of the mixture that can be seen).
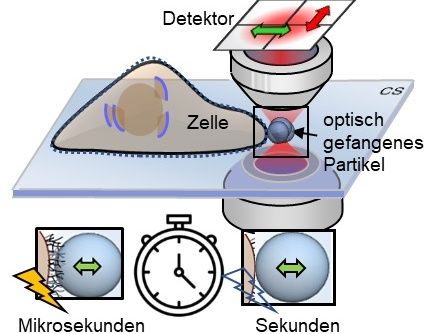
The problem until now has been seeing exactly how long an interaction between a biological molecule and an enzyme takes and how much time elapses between these interactions. This is complicated by the need to distinguish those molecules interacting with the protein and those just passing by. "A freely moving molecule will come in and out of the observation volume very quickly -- on the order of a microsecond. But if it interacts with the enzyme it will sit there for a millisecond," says Levene. "There are three orders of magnitude difference in the length of time that we see this burst of fluorescence. So now it's very easy to discriminate between random occurrences of one ligand and a ligand interacting with the enzyme."
Says Webb: "We see only one fluorescent ligand at a time, so we can now follow the kinetics [movement and behavior] in real time of individual reactions." He adds, "We can actually see the process of interaction."
The creation of the microchip, just 25 millimeters (1 inch) across and containing 25 wells, each with 90,000 tiny holes, has enabled the researchers to increase the number of ligand molecules to naturally occurring molecular concentrations. "To get to these concentrations and still observe one event at a time you need a very small volume. Otherwise you would have all those molecules milling around and we couldn't tell which was interacting with our enzyme and which wasn't," says Levene.
Why observe one molecule at a time instead of a beaker-full, asks Webb. "Because it has become apparent that the behavior of molecules is variable and there are fluctuations in the way they work," he says. "With different molecules and even within individual molecules there is erratic behavior, sometimes pausing, sometimes moving, in milliseconds of time. It is important to understand this movement," he adds, "because molecules that are erratic have less predictable behavior and are likely to be less selective."
Understanding the fluctuations of a single enzyme is particularly important for drug discovery. The Webb and Craighead research groups also are studying the application of their research to advances in DNA sequencing. The sensitivity of the nanofabricated device might permit decoding genetic information by using just one DNA molecule. The technique is compatible on a manufacturing scale and could easily be integrated with other devices, says Korlach. "It is very compact and also very scalable," he notes. This has been demonstrated, he says, by the ability to create millions of waveguides on a single microchip.
Indeed, Webb says, one sequencing run has been limited to 800 base pairs on a strand of DNA. The new technique promises the possibility of decoding continuous sequences of hundreds of thousands of base pairs at a time, he says.
The research was funded by, among others, the U.S. Department of Energy, the National Institutes of Health and the NSF.