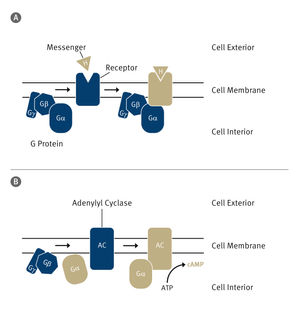
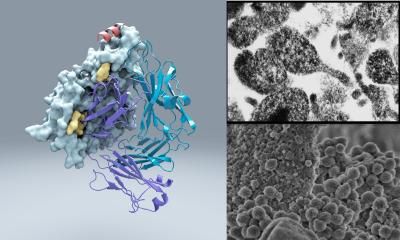
Bacterial protein structure could aid development of new antibiotics
Bacterial cells have an added layer of protection, called the cell wall, that animal cells don't. Assembling this tough armor entails multiple steps, some of which are targeted by antibiotics like penicillin and vancomycin.
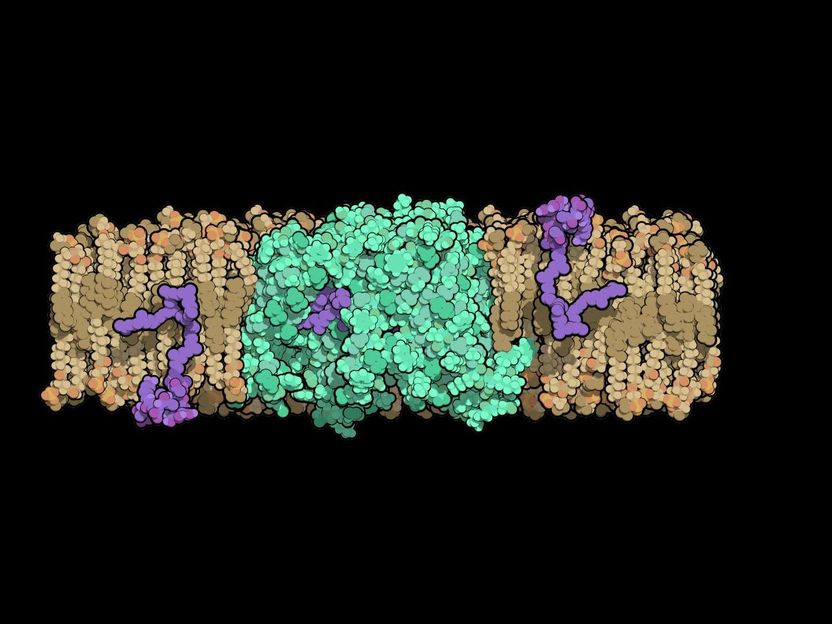
Researchers at Duke University solved the structure of an enzyme that is crucial for helping bacteria build their cell walls. The molecule, called MurJ (shown in green), must flip cell wall precursors (purple) across the bacteria's cell membrane before these molecules can be linked together to form the cell wall. This new structure could be important to help develop new broad-spectrum antibiotics.
Alvin Kuk, Duke University
Yet one step in the process has remained a mystery because the molecular structures of the proteins involved were not known.
Duke University researchers have now provided the first close-up glimpse of a protein, called MurJ, which is crucial for building the bacterial cell wall and protecting it from outside attack.
Antibiotic researchers feel an urgent need to gain a deeper understanding of cell wall construction to develop new antibiotics in the face of mounting antibacterial resistance. In the U.S. alone, an antibiotic-resistant infection called MRSA causes nearly 12,000 deaths per year.
"Until now, MurJ's mechanisms have been somewhat of a 'black box' in the bacterial cell wall synthesis because of technical difficulties studying the protein," said senior author Seok-Yong Lee, Ph.D., associate professor of biochemistry at Duke University School of Medicine. "Our study could provide insight into the development of broad spectrum antibiotics, because nearly every type of bacteria needs this protein's action."
A bacterium's cell wall is composed of a rigid mesh-like material called peptidoglycan. Molecules to make peptidoglycan are manufactured inside the cell and then need to be transported across the cell membrane to build the outer wall.
In 2014, another group of scientists had discovered that MurJ is the transporter protein located in the cell membrane that is responsible for flipping these wall building blocks across the membrane. Without MurJ, peptidoglycan precursors build up inside the cell and the bacterium falls apart.
Many groups have attempted to solve MurJ's structure without success, partly because membrane proteins are notoriously difficult to work with.
In the new study, Lee's team was able to crystallize MurJ and determine its molecular structure to 2-angstrom resolution by an established method called X-ray crystallography --which is difficult to achieve in a membrane protein.
The structure, combined with follow-up experiments in which the scientists mutated specific residues of MurJ, allowed them to propose a model for how it flips peptidoglycan precursors across the membrane.
After determining the first structure of MurJ, Lee's team is now working to capture MurJ in action, possibly by crystallizing the protein while it is bound to a peptidoglycan precursor.
"Getting the structure of MurJ linked to its substrate will be key. It will really help us understand how this transporter works and how to develop an inhibitor targeting this transporter," Lee said.
Lee's group is continuing structure and function studies of other key players in bacterial cell wall biosynthesis as well. Last year, they published the structure of another important enzyme, MraY, bound to the antibacterial muraymycin.