A Protective Cap for Bacterial RNA
Researchers unravel structure and function of bacterial decapping enzyme
For the first time, researchers from Heidelberg University have deciphered the function of the so-called decapping enzyme in bacteria. These molecular helpers remove the protective cap at the start of ribonucleic acid (RNA) molecules. This decapping destabilises the ribonucleic acid, thus allowing degradation to begin in the cells. While these processes are well understood in the messenger RNA of cells in higher organisms, Prof. Dr Andres Jäschke and his bioorganic chemistry working group now revealed these mechanisms in bacterial RNA. Until now, scientists believed that bacteria did not possess this cap structure.
Ribonucleic acids primarily serve as messengers or scaffold molecules in cells, but they also accelerate key biochemical reactions and regulate metabolic processes. In higher organisms, the eukaryotes, messenger RNA (mRNA) usually has a molecular cap at its start; this chemical modification stabilises the messenger RNA, protecting it from degradation and modification. In the prevailing scientific view, bacterial RNA lacks this cap structure. In 2015, however, Prof. Jäschke and his team discovered a modification in certain bacterial RNAs that is structurally similar to the cap on the messenger RNA in eukaryotes.
The cap is nicotinamide adenine dinucleotide (NAD), a coenzyme that plays a key role in metabolism. If NAD is used as a cap in ribonucleic acid, however, it protects the RNA from degradation and modification. Once the NAD cap is removed, the RNA can be degraded in order to initiate metabolic processes. Prof. Jäschke and his team identified an enzyme known as NudC that is responsible for removing the cap. The Heidelberg researchers from the Institute for Pharmacy and Molecular Biotechnology succeeded in analysing NudC from the Escherichia coli bacterium using high-resolution crystal structures, which enabled them to decode the enzyme's function.
Prof. Jäschke emphasises that the structural investigations open up a new field of research, because possible interaction partners of NudC as well as other decapping enzymes in other bacteria need to be identified. The researchers hope their current findings will fuel new interest in identifying unknown cap structures in other microorganisms as well as their functional mechanisms.
Original publication
Most read news
Original publication
K. Höfer, S. Li, F. Abele, J. Frindert, J. Schlotthauer, J. Grawenhoff, J. Du, D.J. Patel and A. Jäschke; "Structure and function of the bacterial decapping enzyme NudC"; Nature Chemical Biology; published online 18 July 2016
Topics
Organizations
Other news from the department science

Get the life science industry in your inbox
By submitting this form you agree that LUMITOS AG will send you the newsletter(s) selected above by email. Your data will not be passed on to third parties. Your data will be stored and processed in accordance with our data protection regulations. LUMITOS may contact you by email for the purpose of advertising or market and opinion surveys. You can revoke your consent at any time without giving reasons to LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin, Germany or by e-mail at revoke@lumitos.com with effect for the future. In addition, each email contains a link to unsubscribe from the corresponding newsletter.
Most read news
More news from our other portals
Last viewed contents
Solipsism_syndrome
Pinolenic_acid
Allopregnanolone
Joseph_von_Gerlach
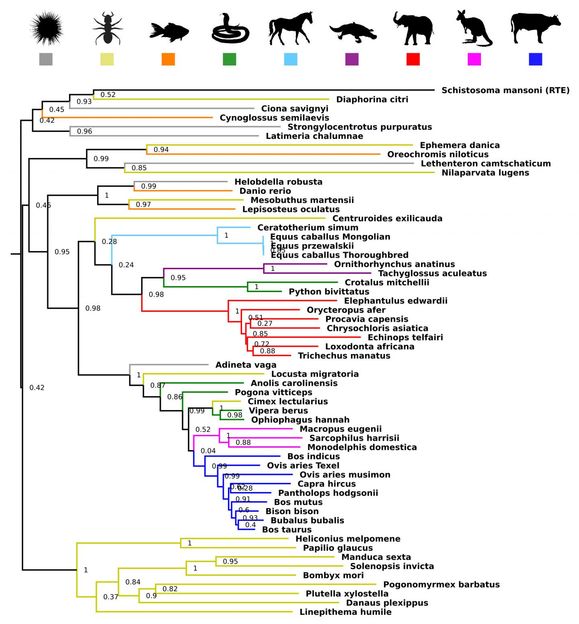
Cross species transfer of genes has driven evolution
Histology