Making sure antibiotics work as they should
A team of ETH Zurich researchers led by professors Nenad Ban and Ruedi Aebersold have studied the highly complex molecular structure of mitoribosomes, which are the ribosomes of mitochondria. Ribosomes are found in the cells of all living organisms. However, higher organisms (eukaryotes), which include fungi, plants, animals and humans, contain much more complex ribosomes than bacteria. In eukaryotes, ribosomes can also be divided into two types: those in the cytosol – which comprises the majority of the cell – and those found in the mitochondria or "power plants" of cells. Mitochondria are only found in eukaryotes.
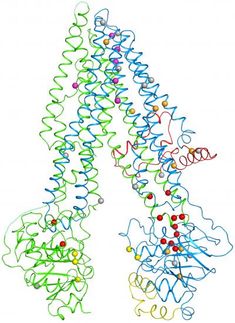
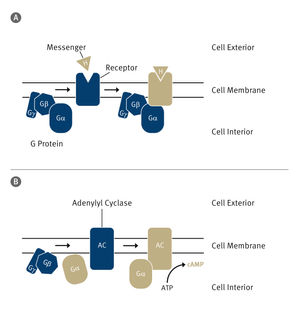
Ribosomes serve as translation devices for the genetic code and produce proteins based on the information stored in DNA. Every ribosome consists of two subunits. The smaller subunit uses transfer ribonucleic acids (transfer RNA or tRNA) to decode the genetic code it receives in the form of messenger RNA, while the larger subunit joins the amino acids delivered by the transfer RNA together like a string of pearls.
Even higher resolution, even more details
Mitochondrial ribosomes are especially difficult to study because they are found only in small amounts and are difficult to isolate. At the beginning of the year, ETH researchers had shed light on the molecular structure of the large subunit of the mitoribosome in mammalian cells to a resolution of 4.9 Å (less than 0.5 nm). However, this resolution was not adequate to reliably build a complete atomic model of this previously unknown structure. The team lead by ETH Professor Nenad Ban has now succeeded in this task and was able to map the entire structure at a resolution of 3.4 Å (0.34 nm). The researchers recently published their findings in the scientific journal Nature.
The scientists used high-resolution cryo-electron microscopy at the Electron Microscopy Center of ETH Zurich (ScopeM) and state-of-the-art mass spectrometry methods in their experiments. Thanks to recent technical advances in cryo-electron microscopy and the development of direct electron detection cameras that can correct for specimen motion during the exposure, it only recently became possible to capture images of biomolecules at a resolution of less than four angstroms.
Improving the effect of antibiotics
In particular, the new images show the details of the peptidyl transferase centre (PTC), which is where the amino acid building blocks are combined. The proteins synthesised in this way then pass through a tunnel, where they finally exit the large subunit of the ribosome.
"This process is medically relevant," said Basil Greber, lead author of the study and postdoctoral researcher in Nenad Ban's group. The reason is that this tunnel is a target for certain antibiotics. The antibiotic becomes lodged in the tunnel and prevents the proteins that have just been synthesized from leaving the tunnel. However, antibiotics should only inhibit protein synthesis in the ribosomes of bacteria.
"For an antibiotic to be used in humans, it must not attack human ribosomes," explains Greber. Antibiotics must inhibit protein synthesis only in bacterial ribosomes. The problem is that mitochondrial ribosomes resemble those of bacteria, which is why certain antibiotics also interfere with mitoribosomes. "This can lead to serious side effects." The ETH researchers' findings will make it possible in the future to design antibiotics that inhibit only bacterial and not mitochondrial ribosomes. This is one basic requirement for using them in clinical applications.
A surprising discovery
The ETH researchers also made an unexpected discovery. They found that mitoribosomes use transfer RNA in two fundamentally different ways. Firstly, the tRNA is used to select the right amino acid for peptide synthesis in the PTC. Secondly, one tRNA is a fixed part of the structure, unlike in all other ribosomes. Although it has been known for quite some time that mitochondrial ribosomes integrated new proteins into their structure over the course of their development, this is the first time that the use of an entirely new RNA molecule was observed. "This demonstrates the great evolutionary plasticity of mitoribosomes," underscored Greber.
The ETH team is now faced with a major, still unresolved task in its research: determining the structure of the smaller subunit of the mitochondrial ribosome. The fact that it is more flexible than the large subunit renders this undertaking an even greater challenge.
Original publication
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic World Mass Spectrometry
Mass spectrometry enables us to detect and identify molecules and reveal their structure. Whether in chemistry, biochemistry or forensics - mass spectrometry opens up unexpected insights into the composition of our world. Immerse yourself in the fascinating world of mass spectrometry!
Topic World Mass Spectrometry
Mass spectrometry enables us to detect and identify molecules and reveal their structure. Whether in chemistry, biochemistry or forensics - mass spectrometry opens up unexpected insights into the composition of our world. Immerse yourself in the fascinating world of mass spectrometry!