Going global
Stowers team reports genome-wide analysis of genes that drive cell division in a multicellular organism
In textbooks, the grand-finale of cell division is the tug-of-war fought inside dividing cells as duplicated pairs of chromosomes get dragged in opposite directions into daughter cells. This process, called mitosis, is visually stunning to observe under a microscope. Equally stunning to cell biologists are the preparatory steps cells take to ensure that the process occurs safely.
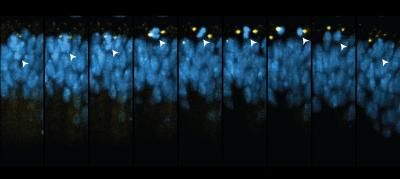
In XZ SPIM time-lapse captures from the developing Drosophila wing disc, mitotic nuclei (His2Av-mRFP, blue) typically rise to the apical epithelial surface, come into contact with MTOCs (GFP-Cnn, yellow), and then move basally after division in a process called interkinetic nuclear migration.
Liang Liang, Ph.D., Stowers Institute for Medical Research
Molecular biologists define those "cell cycle" steps as: G1, when cells survey chromosomes for damage and, if they pass muster, prepare to replicate them; S phase, in which replication occurs; and G2, when cells check duplicated chromosomes one last time for damage and construct the protein machinery required for mitosis.
Failure of a single step is ominous: almost any disease--from autoimmunity to neurodegeneration--is marked, if not caused, by some kind of cycle malfunction. The most obvious is cancer, in which G1 or G2 quality control steps fail and uncontrolled division of tumor cells harboring oncogenic mutations can go totally unchecked.
Halting abnormal cell division thus requires knowing what genes are operational in G1 versus G2. To define them, Stowers Institute of Medical Research Associate Investigator Matt Gibson, Ph.D., undertook a genome-wide comparison of genes expressed during the G1 and G2 phases in larval tissue of the fruitfly Drosophila melanogaster. This work, currently reported online and in the April 14, 2014 print issue of Developmental Cell, catalogues over 300 genes differentially expressed at these steps. Significantly, it is the first to accomplish that in a multicellular organism.
Researchers have previously performed genome-wide screens for cell cycle genes in yeast or cultured animal cells where it is possible to obtain large numbers of cells at precise phases of cell cycle progression. "We started out from the general principle that regulation of the cell cycle in a complex tissue could be very different from how cells divide in a dish," says Gibson. "To understand what happens physiologically, we wanted to apply genome-scale methods to characterize cell division as it occurs in the animal."
Liang Liang, Ph.D., a Gibson lab graduate student, led the all-Stowers team, assisted by Jeff Haug, head of the Cytometry Core, and Genomic Scientist Chris Seidel, Ph.D. Liang began by painstakingly optimizing procedures to dissociate wing disc tissue but keep it viable long enough to analyze. Haug, who worked with her in this phase of the project, calls this a feat in itself, requiring what he calls "institutional memory" of methods used by other Stowers investigators to prepare cells from Drosophila tissues.
The team then stained living target cells with dye that labels DNA. That procedure allowed them to sort cells into two bins using a technique called flow cytometry: one bin contained cells with one copy of the genome (cells in G1) and the other contained cells with two genomes (those in G2, which had replicated their DNA but not yet divided).
Microarray analysis then identified every gene exhibiting different levels of expression in the G1 or G2 cell populations. Some were expressed in both, but 431 genes were upregulated in G1 and 336 during G2. The team further validated candidates by genetically "knocking down" each separately in wing discs and examining adult fly wings for defects, which they saw in 80% of cases.
Gibson says that many usual suspects were activated at the "right" time. "Things that control DNA replication were enriched in G1, while factors regulating mitosis were expressed in G2," he explains. The surprise came when the team compared data with parallel profiles obtained using cultured Drosophila S2 cells: many of the roughly 200 genes differentially expressed in G1 or G2 in disc cells were uniformly expressed in S2 cells. Likewise, close to 100 genes differentially expressed in S2 cells showed uniform expression in disc cells.
"What was exciting was the plasticity we saw in cell cycle regulation of gene expression," says Liang, noting different profiles seen in disc versus S2 cells. "Every animal uses the same cell cycle machinery, but that machinery may be regulated very differently depending on the cell type, even in the same organism."
First and foremost, the work provides a searchable resource freely available to scientists and citizens alike at the Stowers Original Data Repository. There, you can see if your favorite gene is expressed in G1 or G2 in wing discs or S2 cells and view pictures of what fly wings look like when that gene is deficient.
The work also identifies brand-new suspects for the Gibson lab. Wing disc cells and cells that line almost every mammalian body cavity are epithelial cells, which grow in sheets. They divide in a peculiar fashion: right before mitosis the nucleus of an elongated epithelial cell moves into one end of the cell for cell division, a process called interkinetic nuclear movement (IKNM). Gibson is interested in the mechanics of IKNM, in part because epithelial cell cancers, termed carcinomas, comprise over 80% of all malignancies. "One motivation for this study was to discover what links IKNM to the cell cycle," he says. "That required a global view."
He was right. The paper reports two genes that when knocked down in wing discs disrupt IKNM as cells divide without interrupting division itself. Intriguingly, one is a long noncoding RNA (lncRNA), a currently mysterious class of RNAs that does not encode proteins but instead may regulate downstream gene expression.
Seidel, who helped analyze data for the paper, says next-gen technologies free researchers to ask unbiased questions, without which the team would have surely missed the lncRNA. He compares "old" versus "new" genome exploration to mapping the earth before and after satellites. "Before, you sailed from place to place for hundreds of years establishing landmarks to create a map," he says. "Afterwards, a few hours' worth of data collection offered a comprehensive, global view."
Gibson concurs, but with a nod to the field's Magellans and Ponce de Leons. "Historically, scientists studied cell cycle control by taking a gene-by-gene and protein-by-protein approach, usually in cultured cells," he says. "Those pioneers provided immense insight into how cell division works. Now we have tools to determine how that fundamental process is fine-tuned to operate in the complex and varied contexts present in a multicellular animal."