Epigenetics enigma resolved
First structure of enzyme that removes methylation
Scientists have obtained the first detailed molecular structure of a member of the Tet family of enzymes.
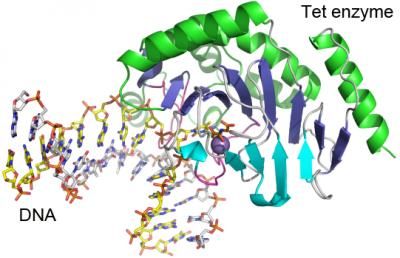
This is the structure of the Tet enzyme with DNA. Note the purple ball at the active site, close to which one DNA base is flipped out of the double helix. Also note the degree to which the double helix is bent.
Xiaodong Cheng, Emory University
The finding is important for the field of epigenetics because Tet enzymes chemically modify DNA, changing signposts that tell the cell's machinery "this gene is shut off" into other signs that say "ready for a change."
Tet enzymes' roles have come to light only in the last five years; they are needed for stem cells to maintain their multipotent state, and are involved in early embryonic and brain development and in cancer.
The results, which could help scientists understand how Tet enzymes are regulated and look for drugs that manipulate them, are scheduled for publication in Nature.
Researchers led by Xiaodong Cheng, PhD, determined the structure of a Tet family member from Naegleria gruberi by X-ray crystallography. The structure shows how the enzyme interacts with its target DNA, bending the double helix and flipping out the base that is to be modified.
"This base flipping mechanism is also used by other enzymes that modify and repair DNA, but we can see from the structure that the Tet family enzymes interact with the DNA in a distinct way," Cheng says.
Cheng is professor of biochemistry at Emory University School of Medicine and a Georgia Research Alliance Eminent Scholar. The first author of the paper is research associate Hideharu Hashimoto, PhD. A team led by Yu Zheng, PhD, a senior research scientist at New England Biolabs, contributed to the paper by analyzing the enzymatic activity of Tet using liquid chromatography–mass spectrometry.
Using oxygen, Tet enzymes change 5-methylcytosine into 5-hydroxymethylcytosine and other oxidized forms of methylcytosine. 5-methylcytosine (5-mC) and 5-hydroxymethylcytosine (5-hmC) are both epigenetic modifications of DNA, which change how DNA is regulated without altering the letters of the genetic code itself.
5-mC is generally found on genes that are turned off or on repetitive regions of the genome. 5-mC helps shut off genes that aren't supposed to be turned on (depending on the cell type) and changes in 5-mC's distribution underpin a healthy cell's transformation into a cancer cell.
In contrast to 5-mC, 5-hmC appears to be enriched on active genes, especially in brain cells. Having a Tet enzyme form 5-hmC seems to be a way for cells to erase or at least modify the "off" signal provided by 5-mC, although the functions of 5-hmC are an active topic of investigation, Cheng says.
Alterations of the Tet enzymes have been found in forms of leukemia, so having information on the enzymes' molecular structure could help scientists design drugs that interfere with them.
N. gruberi is a single-celled organism found in soil or fresh water that can take the form of an amoeba or a flagellate; its close relative N. fowleri can cause deadly brain infections. Cheng says his team chose to study the enzyme from Naegleria because it was smaller and simpler and thus easier to crystallize than mammalian forms of the enzyme, yet still resembles mammalian forms in protein sequence.
Mammalian Tet enzymes appear to have an additional regulatory domain that the Naegleria forms do not; understanding how that domain works will be a new puzzle opened up by having the Naegleria structure, Cheng says.
Other news from the department science

Get the life science industry in your inbox
By submitting this form you agree that LUMITOS AG will send you the newsletter(s) selected above by email. Your data will not be passed on to third parties. Your data will be stored and processed in accordance with our data protection regulations. LUMITOS may contact you by email for the purpose of advertising or market and opinion surveys. You can revoke your consent at any time without giving reasons to LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin, Germany or by e-mail at revoke@lumitos.com with effect for the future. In addition, each email contains a link to unsubscribe from the corresponding newsletter.
Most read news
More news from our other portals
Last viewed contents
Stomach_cancer_in_cats_and_dogs
Intersex_surgery
Biotest starts combination therapy study with BT-062 in multiple myeloma
Arrhythmogenic_right_ventricular_dysplasia
Amplification_(psychology)
ChemMaps lets researchers navigate the chemical universe
American_Synesthesia_Association
Mary_Ainsworth
Category:Tetracyclic_antidepressants
-phob-
Cryo-force spectroscopy reveals the mechanical properties of DNA components