Algorithms for identifying new "cancer genes"
Advertisement
It is estimated that the number of cancer cases worldwide will double by 2040. This makes the search for genes that cause cancer even more important. A team of researchers from the University of Bern and Inselspital, University Hospital Bern, has now developed algorithms that massively simplify the hunt for "cancer genes" in a poorly understood part of our genome.
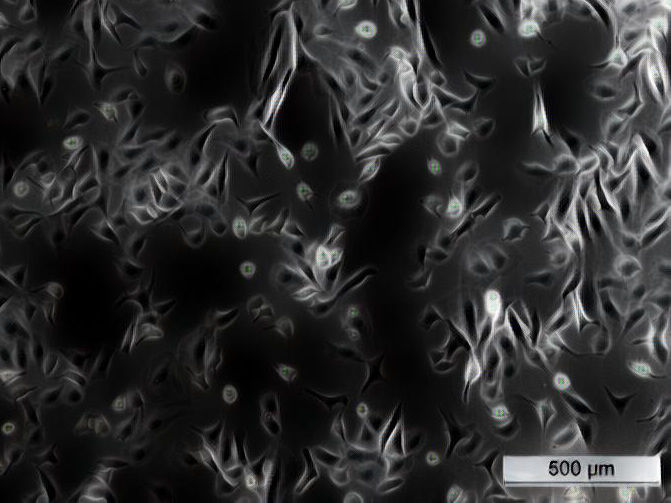

Microscopy image of a cell line used to identify long noncoding RNAs related to lung cancer.
© Taisia Polidori, Universität Bern
Cancer is caused by mutations in the genome of cells. Mutated cells grow in an uncontrolled way, adapt to new conditions and can escape the body's defence mechanisms. For this reason, researchers are increasingly focusing on the genetics of tumors. Looking at the genetic profile of these malignant cells helps us to understand how a cancer develops and what drives its spread. This also can provide clues for therapeutic targets. The hunt for mutated genes, which cause cancer – so called "driver genes", is made possible through the latest technologies for DNA sequencing.
The driver genes in tumors are identified based on their patterns of genetic mutations by means of sophisticated algorithms. Such a sensitive method must be carefully calibrated. "Think of the weighing scale in your home, which must be adjusted from time to time to show the correct weight. In a similar way, methods to search for driver-genes must be calibrated using "benchmarks", that is, sets of already-known cancer genes", says Rory Johnson. He conducts his research at the Department for BioMedical Research of the University of Bern (DBMR) and the Inselspital, University Hospital Bern and is a member of the National Center of Competence in Research RNA & Disease. His group has assembled a dataset of genes, which significantly facilitates researchers’ search for novel cancer driver genes.
Cancer genes in the "dark matter" of our genome
The term "dark matter" of our genome refers to the over 95 % of it, which does not contain instructions for building proteins. Numerous studies indicate that a part of this "dark matter", called long non-coding RNAs or "lncRNAs", play important roles in tumorigenesis and cancer progression. If we consider DNA (deoxyribonucleic acid) to be the fixed blueprint for an organism, then RNA (ribonucleic acid) represents a "real-time" readout of that blueprint that dynamically changes in response to the needs of the cell and organism. The biological roles and molecular mechanisms of just a tiny fraction of these lncRNAs have been studied to date. "Cancer lncRNAs that provoke tumours represent an exciting new focus for the development of cancer therapies", says Andrés Lanzos, first author of the study from the DBMR and Inselspital, University Hospital Bern, and NCCR RNA&Disease.
Cancer researchers have traditionally focused their efforts on the approximately 19,000 "classical" protein coding genes in the human genome. For these genes, a benchmark has long existed, consisting of genes known to play roles in tumorigenesis and cancer development. The team led by Rory Johnson is focused on searching for cancer lncRNAs using maps of tumour mutations from the International Cancer Genome Consortium. For this, the researchers have developed statistical methods to identify cancer lncRNAs. They wanted to calibrate the accuracy of these new methods with the help of a benchmark like is the case for classical protein-coding genes. For this purpose, the team assembled a list of 122 long-non coding RNAs that have been implicated in cancer with high confidence.
High quality predictions possible
"This dataset of 122 cancer-lncRNAs has already proved an invaluable resource in many ways", says Johnson. The team has used it to calibrate their algorithms for cancer lncRNA discovery, and it has already demonstrated that such algorithms make high quality predictions, including dozens of completely new cancer lncRNAs. Their "ExInAtor" algorithm has already been successfully used for the efforts of the International Cancer Genome Consortium, which has just published their results in a series of papers in the journal Nature and elsewhere. This large-scale project also involved Mark Rubin, Director of the Department for BioMedical Research (DBMR) of the University of Bern and the Inselpital, University Hospital Bern.
"We are convinced that this gene dataset proves a unique resource to better understand the properties of this poorly-understood class of lncRNA genes", explains Johnson. "On the one hand this should help researchers to refine their methods for searching for cancer lncRNAs, so that we can extend the list of cancer lncRNAs, and on the other hand we hope that this enables the development of a new generation of personalized therapies for cancer patients", he adds.
Original publication
Joana Carlevaro-Fita, Andrés Lanzós, Lars Feuerbach, Chen Hong, David Mas-Ponte, Jakob Skou Pedersen, PCAWG Drivers and Functional Interpretation Group, Rory Johnson & PCAWG Consortium; "Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis"; Communications Biology; 5 February 2020