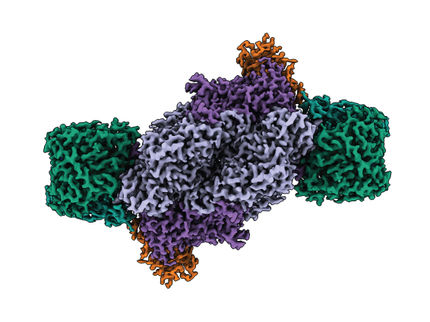
Structure of protein nano turbine revealed
Scientists determine the first structure of a cell’s rotary engine using state-of-art microscopy
Cells rely on protein complexes known as ATP synthases or ATPases for their energy needs – adenosine triphosphate (ATP) molecules power most of the processes sustaining life. Structural biologist Professor Leonid Sazanov and his research group from the Institute of Science and Technology Austria (IST Austria) in Klosterneuburg, Austria have now determined the first atomic structure of the representative of the V/A-ATPase family, filling in the gap in the evolutionary tree of these essential molecular machines. These results obtained using the latest cryo-electron microscopy methods revealed a turbine or water mill similar structure of the enzyme.
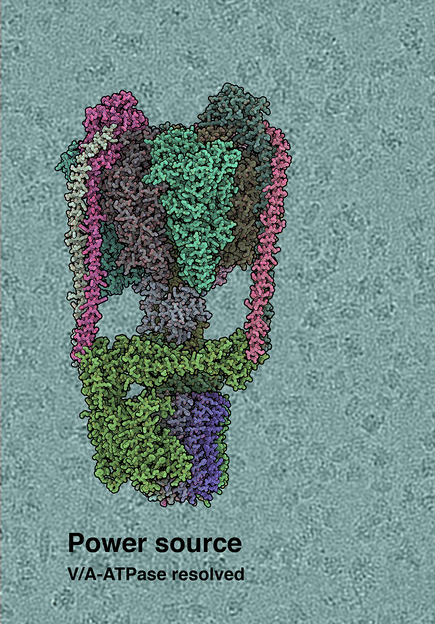
Cryo-EM structure of the T. thermophilus V/A-type ATP synthase. The background shows raw cryo-EM micrograph, with individual ATPase molecules visible.
(c) by IST Austria, 2019
Rotary power
ATP synthases/ATPases are large membrane protein complexes which share overall gross building plans and rotary catalysis mechanisms. This protein family includes F-type enzyme found in mitochondria (power factories of the cell), chloroplasts (organelles that conduct photosynthesis in plants) and bacteria; V (vacuolar)-type found in intracellular compartments in eukaryotes (higher organisms with a nucleus) and A (archaeal)-type found in prokaryotes - archaea (ancient microorganisms) and some bacteria. The proteins consist of a globular F1/V1/A1 domain synthesizing/hydrolyzing ATP and a membrane-embedded Fo/Vo/Ao domain translocating protons. F-type enzymes are well studied and their atomic structures are known. F1 and Fo domains are connected by the central stalk or shaft/rotor, which rotates inside F1 driving ATP synthesis. The rotor is attached to the so-called c-ring which is driven by the proton flow between the ring and the membrane-embedded stator. The rotation of F1 itself is prevented by the single peripheral stalk, which keeps F1 anchored to the stator. The overall architecture of this unique molecular machine thus resembles that of a turbine or a water mill, driven by the flow of protons rather than water.
Different flavors of ATPases
F- and A-type enzymes usually function to produce ATP, driven by proton flow across the membrane. V-type enzymes usually work in reverse, using ATP to pump protons. V- and A-ATPases are similar structurally but they differ from the F-type by having two or three peripheral stalks and additional connecting protein subunits between V1 and Vo. V-type enzymes probably evolved from the A-type and because of these similarities A-type is also termed V/A-ATPase. Some bacteria, including Thermus thermophilus, acquired an A-type enzyme. Long Zhou, postdoc in the Sazanov research group of IST Austria, has purified and studied this enzyme (ThV1Vo) by cryo-EM. In contrast to F-type, for V-type ATPases only the structures of the isolated V1 and Vo domains were determined previously. How V1 is coupled to Vo was therefore not known, and the knowledge about the full catalytic cycle was lacking.
Plasticity and competition
The scientists determined not one, but in total five structures of the entire ThV1Vo enzyme, using cryo-electron microscopy methods developed recently in the so-called “resolution revolution” of this technique. The structures represent several conformational states of the enzyme differing by the position of the rotor inside the stator. Global conformational plasticity of ThV1Vo is revealed as substantial V1 wobbling in space in transition from one state to another. It is a result of mechanical competition between rotation of the bent central rotor and stiffness of the stator. V1-Vo coupling is achieved via close structural and electrostatic match between the shaft and V-type specific subunit linking it to the c-ring. The visualization of the proton path revealed significant differences in the distribution of charged protein residues from that in F-ATPases, with a stricter “check-point” preventing “slipping” of the enzyme.
Why additional complexity?
Instead of a single peripheral stalk of F-type enzymes, A-types such as ThV1Vo have two peripheral stalks, while eukaryotic V-types have three. But what is the advantage of the additional complexity in the already very large protein assembly, along with additional subunits linking V1 and Vo? The F1/V1 domain has a three-fold symmetry and so one ATP molecule is produced (or consumed) per each 120° rotation of the stator inside F1/V1. Professor Leonid Sazanov says: “In V/A-ATPases this step is a one-off 120° rotation, in contrast to F-ATP synthase where it is divided into several sub-steps. Thus, greater plasticity may be required in ThV1Vo in order to link these 120° steps in V1 to smaller per c subunit steps in the Vo c12-ring. This additional flexibility may be afforded in V-types by the additional peripheral stalks and connecting subunits. Our new structures show how this is achieved, providing a framework for the entire V-ATPase family”.
Original publication
Zhou, L. and Sazanov, L.A.; "Structure and Conformational Plasticity of the Intact Thermus thermophilus V/A-type ATPase"; Science; 2019, in press.